Exploration of potential inhibitors of poxin protein in monkeypox virus through molecular docking techniques: An in-silico drug repurposing study
Authors
- K. M. Ferdousul HaqueDepartment of Pharmaceutical Chemistry, University of Dhaka, Dhaka, Bangladesh
- Md. Khalid HossainDepartment of Pharmaceutical Chemistry, University of Dhaka, Dhaka, Bangladesh
- Md. Shafiul HossenDepartment of Pharmacy, State University of Bangladesh, Dhaka, Bangladesh
- Mohammad Sharifur RahmanDepartment of Pharmaceutical Chemistry, University of Dhaka, Dhaka, Bangladesh
- Mohammad A. RashidDepartment of Pharmaceutical Chemistry, University of Dhaka, Dhaka, Bangladesh
DOI:
Keywords
Downloads
Correspondence
Publication history
Responsible editor
Reviewers
Funding
Ethical approval
Trial registration number
Copyright
Published by Bangladesh Medical University (former Bangabandhu Sheikh Mujib Medical University).
Background: Monkeypox virus (MPXV) has re-emerged as a major public health concern due to its expanding global spread. However, limited therapeutic options hinder disease control. Molecular docking offers a valuable computational approach for identifying potential antiviral candidates.
Methods: This study docked several antiviral drugs, including tecovirimat, tipranavir, remdesivir, fluocinolone, molnupiravir, famciclovir, acyclovir, cidofovir, and brincidofovir, against the monkeypox poxin protein (Protein Data Bank Identification (PDB ID): 8C9K). These drugs were selected based on reported anti-orthopoxvirus activity, clinical availability, and frequent prescription. Drug-likeness screening identified promising inhibitors. Molecular docking was performed using PyRx v0.9.8 with a 25 × 25 × 25 Å grid centered on the active site (x = 28.7, y = 56.9, z = 37.3). Tecovirimat served as the reference drug. Adverse drug monitoring event (ADME) and toxicity predictions assessed topological polar surface area, lipophilicity, solubility, bioavailability, blood–brain barrier permeability, P-glycoprotein interaction, median lethal dose (LD₅₀), and toxicity class by the Swiss ADME webtool.
Results: Fluocinolone (−8.8 kcal/mol; Kilocalorie per mole), remdesivir (−8.9 kcal/mol), and tipranavir (−9.6 kcal/mol) showed stronger binding affinities than tecovirimat (−7.6 kcal/mol), while molnupiravir exhibited comparable affinity (−7.3 kcal/mol). The pharmacokinetic and toxicity profiles of fluocinolone, molnupiravir, and tipranavir were almost similar to the reference drug. Remdesivir showed minor physicochemical differences but was predicted to be safe.
Conclusion: Tipranavir, remdesivir, fluocinolone, and molnupiravir emerged as promising MPXV poxin protein inhibitors, requiring further experimental validation.
Human monkeypox (HMPX) is a zoonotic infection caused by the monkeypox virus (MPXV), a double-stranded DNA virus belonging to the Orthopoxvirus genus within the Poxviridae family [1]. Since the global eradication of smallpox in 1980, MPXV has emerged as the most significant orthopoxvirus to public health worldwide [2]. The first human case was documented in 1970 in the Democratic Republic of the Congo in an unvaccinated child, and the virus subsequently established endemicity in Central and West Africa [1, 3]. In May 2022, an unprecedented multinational outbreak across non-endemic regions with no clear epidemiological links marked a pivotal moment, refocusing global attention on MPXV as an emerging pathogen of international concern [1, 4].
Transmission primarily occurs through direct contact with infectious skin lesions, body fluids, or contaminated materials, as well as via respiratory droplets during prolonged close contact [5]. Following entry through broken skin or mucous membranes, the virus replicates locally before disseminating to regional lymph nodes, with an incubation period typically ranging from 7 to 14 days [1, 4, 6]. While historical outbreaks were often limited in scale and geography, the 2022 epidemic highlighted the virus capacity for rapid, widespread human-to-human transmission beyond endemic regions, raising significant concerns about its pandemic potential [7]. Disease severity and case fatality rates exhibit considerable geographical and demographic variation, with estimates reaching up to 10% in certain endemic settings, particularly among unvaccinated populations [8].
The discontinuation of routine smallpox vaccination following its eradication has led to waning population immunity, increasing global susceptibility to MPXV and sparking debate on the need for targeted vaccination strategies in at-risk groups [9, 10]. Current management of monkeypox is largely supportive, as no antiviral therapy has been conclusively proven effective in human clinical trials [7]. This therapeutic gap underscores the urgent need for the development of targeted antiviral agents.
A promising strategy involves disrupting critical viral proteins essential for pathogenesis and immune evasion [11]. Among these, poxin, a viral nuclease that plays a key role in subverting host innate immunity [12]. Poxin specifically cleaves 2',3'-cyclic guanosine monophosphate–adenosine monophosphate (cGAMP), a crucial second messenger molecule produced upon cytosolic DNA sensing [13, 14]. cGAMP activates the STING (Stimulator of Interferon Genes) pathway, triggering a robust interferon-mediated antiviral response. By degrading cGAMP, poxin effectively disables this host defense mechanism, allowing the virus to replicate unchecked. Inhibiting poxin function, therefore, represents a rational strategy to restore host immunity and curb viral replication [15, 16].
Molecular docking is a foundational computational technique in structure-based drug design that predicts the preferred orientation and binding affinity of a small molecule (ligand) within a protein's active site [17]. By simulating these interactions, docking facilitates the virtual screening of compound libraries, aids in lead optimization, and provides mechanistic insights into ligand-target binding [18, 19]. This approach is particularly valuable for drug repurposing, where existing, well-characterized drugs can be rapidly evaluated against new targets.
Given the absence of approved treatments and the validated role of poxin in MPXV immune evasion, this study employed molecular docking to computationally screen and evaluate a panel of existing antiviral drugs as potential inhibitors of the monkeypox poxin protein, with the goal of identifying promising candidates for further experimental investigation.
Selection and preparation of ligands
Antiviral drugs active against the orthopoxvirus were included in the study for docking. We identified eight drugs for the study based on a rigorous literature review due to their antiviral activity against the orthopoxvirus. Furthermore, the drugs are available in our local market and are frequently prescribed as an antiviral agent. According to the structure of the drugs was retrieved from the online database PubChem (https://pubchem.ncbi.nlm.nih.gov/) and analyzed for three dimensional (3D) conformations into the drug discovery platform. The 3D conformers of the selected ligands were downloaded in structure data file (SDF) format [20].
Preparation of target protein
The crystal structure of the poxin protein (PDB ID: 8C9K) of the monkeypox virus was retrieved from the Research Collaboratory for Structural Bioinformatics (RCSB) Protein Data Bank (PDB) database (https://www.rcsb.org/) in PDB format [21, 22]. The downloaded structure had a resolution of 1.72 Angstrom (Å). Using PyMOL (v2.5.2) software (Schrodinger, a Limited Liability Company), the protein structure was cleaned by removing unnecessary atoms and molecules, including ligands [23]. The receptor-binding domain (RBD) of the poxin protein was determined in the crystal structure, and the excess protein chains were removed using PyMOL. The protein chains were saved in PDB format for molecular docking.
Molecular docking
Molecular docking of the selected ligands (tecovirimat, tipranavir, remdesivir, fluocinolone, molnupiravir, famciclovir, acyclovir, cidofovir, brincidofovir) against the targeted monkeypox poxin protein (8C9K) were performed using the PyRx v0.9.8 programme. Before that, energy minimization of the structures was performed using Avogadro v1.0 software using auto-optimization tool (Force field: Static variant of Merck Molecular Force Field,1994 (MMFF94s); 4 steps per gradient; conjugate descent algorithm). The docking protocol was validated by re-docking the native ligand, yielding a root mean square deviation (RMSD) of <2.0 Å. The grid box was centered on the active site residues (x=28.7, y=56.9, z=37.3) with dimensions 25×25×25 Å. Exhaustiveness was set to 8 for all docking runs [24]. For every protein-ligand combination, the Dassault Systèmes' BIOVIA Discovery Studio 2021 Client version 21.1.0 software was used to analyze the binding affinity (kcal/mol), noncovalent interactions, and docking orientations. [25]. BIOVIA Discovery Studio provided the two dimensional (2D), and 3D schematic sketches of the protein-ligand docking complexes [26].
Adverse drug monitoring event and toxicity prediction
According to Daina et al. [27] and Kim et al. [28], the SwissADME website (https://www.swissadme.ch/) incorporates the canonical simplified molecular input line entry system (SMILES) of the top-docking ligands that were extracted from the public chemical (PubChem) database [28]. Each ligand's ADME (absorption, distribution, metabolism, and excretion) statistics were supplied by SwissADME. After that, the toxicity profile of each ligand was estimated using the ProTox- service (https://tox.charite.de/protox3) [29]. The physicochemical, pharmacokinetic, and pharmacodynamic properties of every ligand were documented using these two sources. ADME and toxicity prediction process examined the topological polar surface area (TPSA), lipophilicity (MLogP), water solubility (LogS), bioavailability score etc. BOILED-Egg model and bioavailability radar were used to evaluate gastrointestinal absorption (GI) module, blood–brain barrier permeability (BBB), and interaction with P -glycoprotein (P -gp) substrate.


The molecular docking analysis of selected antiviral compounds against the target poxin protein (PDB: 8C9K) revealed substantial variation in predicted binding affinities as shown in Table 1. Among the tested ligands, tipranavir exhibited the most favorable interaction with a binding energy of -9.6 kcal/mol, significantly surpassing the reference drug tecovirimat (-7.6 kcal/mol). This complex was stabilized by approximately 15 interactions with active site residues. Remdesivir (-8.9 kcal/mol) and fluocinolone (-8.8 kcal/mol) also demonstrated stronger theoretical binding than the reference. The affinity of molnupiravir (-7.3 kcal/mol) was closely comparable to tecovirimat, whereas famciclovir, acyclovir, cidofovir, and brincidofovir exhibited comparatively weaker binding, with energies ranging from -5.8 to -6.1 kcal/mol. The detailed two-dimensional (2D), and three-dimensional (3D) interaction diagrams that illustrate hydrogen bonds and hydrophobic interactions between the ligands and the active-site residues of the poxin protein are presented in Supplementary file (Figure 3).
Table 1 The noncovalent (hydrogen bonding and hydrophobic) interactions and binding affinities between the test drugs and the reference inhibitor (tecovirimat)
Sample name | Binding affinity (Kcal/ mol) | Interactions with amino acids a |
Tecovirimat | -7.6 | Lys 42, Leu 84, Asp 83, Val 35, Ser 34 |
Tipranavir | -9.6 | Gly 63, Ser 109, Ala 64, Pro 20, Gly 104, Phe 19, Ala 18, Asp 111, Arg 184, Arg 60, Ala 145, Ile 141, Arg 182, Ala 129, Tyr 138 |
Remdesivir | -8.9 | Ala 145, Lys 59, Lys 178, Cys 113, Asp 111, Ile 180, Glu 179, Arg 60, Lys 146 |
Fluocinolone | -8.8 | Glu 179, Arg 60, Lys 178, Ala 145, Lys 142 |
Molnupiravir | -7.3 | Lys 59, Lys 178, Asn 149, Tyr 131, Glu 179, Arg 60, Asn 58 |
Famciclovir | -6.1 | His 122, Gln 76, Lys 133, Tyr 74, Tyr 39, Lys 41 |
Acyclovir | -6.1 | Asp 111, Arg 184, Ala 64, Lys 186 |
Cidofovir | -5.9 | Arg 182, Glu 179, Arg 60, Asp 111, Tyr 131, Ala 145 |
Brincidofovir | -5.8 | Lys 178, Asp 111, Tyr 131, Val 130, Ala 145, Tyr 138, Asn 149, Lys 59, Glu 179 |
a Interactions are classified majorly as hydrogen bonds (H-bond), hydrophobic (Hyd), and electrostatic (Elec) bonds. | ||
Evaluation of ADME and physicochemical properties indicated that all candidate molecules possessed a higher topological polar surface area (TPSA) relative to tecovirimat (Table 2). Lipophilicity, as measured by MLogP, was within the acceptable threshold (<5) for all compounds, consistent with Lipinski’s Rule of Five. Aqueous solubility, classified per the estimated solubility (ESOL) method, ranged from the high solubility of cidofovir to the poor solubility of tipranavir. Predicted human gastrointestinal absorption was high for fluocinolone, famciclovir, and acyclovir, mirroring the reference drug, but was low for the remaining candidates. Bioavailability scores were largely similar to tecovirimat (0.55), with notable exceptions being the lower scores for remdesivir (0.17) and cidofovir (0.11). Fluocinolone, remdesivir, tipranavir, and famciclovir were predicted substrates for P-glycoprotein.
Table 2 Adverse drug monitoring event properties of the tested compounds and reference drug tecovirimat
Property | Tecovirimat | Tipranavir | Remdesivir | Fluocinolone | Molnupiravir | Famciclovir | Acyclovir | Cidofovir | Brincidofovir |
MLogP | 3.50 | 3.74 | 0.18 | 0.70 | -0.75 | -0.15 | -1.83 | -2.66 | 1.63 |
TPSA (Ų) | 66.48 | 113.97 | 213.36 | 115.06 | 143.14 | 122.22 | 119.05 | 157.71 | 155.94 |
MW (g/mol) | 376.33 | 602.66 | 602.58 | 412.42 | 329.31 | 321.33 | 225.20 | 279.19 | 561.69 |
ESOL Class (Solubility) | Soluble | Poorly | Moderately | Soluble | Very high | Very high | Very high | Highly | Moderately |
LogS (ESOL) | -3.72 | -7.49 | -4.12 | -2.95 | -0.83 | -1.49 | -0.41 | 0.85 | -4.80 |
Bioavailability score | 0.55 | 0.56 | 0.17 | 0.55 | 0.55 | 0.55 | 0.55 | 0.11 | 0.55 |
Lipinski’s RO5 violation | 0 | 1 | 2 | 0 | 0 | 0 | 0 | 0 | 1 |
GI absorption | High | Low | Low | High | Low | High | High | Low | Low |
BBB permeation | Yes | No | No | No | No | No | No | No | No |
P-gp substrate | No | Yes | Yes | Yes | No | Yes | No | No | No |
LD50 (mg/kg) | 2028 | 300 | 1000 | 3100 | 826 | 570 | 5000 | 1681 | 5010 |
Ghose filter violation | 0 | 4 | 3 | 0 | 1 | 0 | 1 | 1 | 3 |
MLogP indicate lipophilicity; TPSA, Topological polar surface area; MW, molecular weight; LogS, solubility in water; ESOL, water solubility course; RO5, rule of five; GI, gastrointestinal absorption; BBB, blood-brain barrier permeability; P-gp, P-glycoprotein; LD50, Lethal dosage50 | |||||||||
Toxicological profiling assigned compounds to various acute oral toxicity classes. Remdesivir, molnupiravir, famciclovir, and cidofovir were categorized in class 4; tipranavir in class 3; and brincidofovir in class 6. Fluocinolone and acyclovir shared the same class 5 designation as tecovirimat, suggesting a similar safety margin at the administered dose. The majority of candidates were predicted to be inactive or to present only moderate risks for hepatotoxicity, carcinogenicity, mutagenicity, and cytotoxicity endpoints. Notably, remdesivir, cidofovir, and brincidofovir were consistently flagged as inactive or moderate across all evaluated toxicity profiles, indicating a potentially favorable safety window (Table 3).
Variables | Frequency (%) |
Indication of colposcopy |
|
Visual inspection of the cervix with acetic acid positive | 200 (66.7) |
Abnormal pap test | 13 (4.3) |
Human papilloma virus DNA positive | 4 (1.3) |
Suspicious looking cervix | 14 (4.7) |
Others (per vaginal discharge, post-coital bleeding) | 69 (23.0) |
Histopathological diagnosis | |
Cervical Intraepithelial Neoplasia 1 | 193 (64.3) |
Cervical Intraepithelial Neoplasia 2 | 26 (8.7) |
Cervical Intraepithelial Neoplasia 3 | 32 (10.7) |
Invasive cervical cancer | 27 (9.0) |
Chronic cervicitis | 17 (5.6) |
Squamous metaplasia | 5 (1.7) |
Groups based on pre-test marks | Pretest | Posttest Marks (%) | Difference in pre and post-test marks (mean improvement) | P |
Didactic lecture classes | ||||
<50% | 36.6 (4.8) | 63.2 (9.4) | 26.6 | <0.001 |
≥50% | 52.8 (4.5) | 72.4 (14.9) | 19.6 | <0.001 |
Flipped classes | ||||
<50% | 36.9 (4.7) | 82.2 (10.8) | 45.4 | <0.001 |
≥50% | 52.8 (4.6) | 84.2 (10.3) | 31.4 | <0.001 |
Data presented as mean (standard deviation) | ||||
Background characteristics | Number (%) |
Age at presentation (weeks)a | 14.3 (9.2) |
Gestational age at birth (weeks)a | 37.5 (2.8) |
Birth weight (grams)a | 2,975.0 (825.0) |
Sex |
|
Male | 82 (41) |
Female | 118 (59) |
Affected side |
|
Right | 140 (70) |
Left | 54 (27) |
Bilateral | 6 (3) |
Delivery type |
|
Normal vaginal delivery | 152 (76) |
Instrumental delivery | 40 (20) |
Cesarean section | 8 (4) |
Place of delivery |
|
Home delivery by traditional birth attendant | 30 (15) |
Hospital delivery by midwife | 120 (60) |
Hospital delivery by doctor | 50 (25) |
Prolonged labor | 136 (68) |
Presentation |
|
Cephalic | 144 (72) |
Breech | 40 (20) |
Transverse | 16 (8) |
Shoulder dystocia | 136 (68) |
Maternal diabetes | 40 (20) |
Maternal age (years)a | 27.5 (6.8) |
Parity of mother |
|
Primipara | 156 (78) |
Multipara | 156 (78) |
aMean (standard deviation), all others are n (%) | |
Background characteristics | Number (%) |
Age at presentation (weeks)a | 14.3 (9.2) |
Gestational age at birth (weeks)a | 37.5 (2.8) |
Birth weight (grams)a | 2,975.0 (825.0) |
Sex |
|
Male | 82 (41) |
Female | 118 (59) |
Affected side |
|
Right | 140 (70) |
Left | 54 (27) |
Bilateral | 6 (3) |
Delivery type |
|
Normal vaginal delivery | 152 (76) |
Instrumental delivery | 40 (20) |
Cesarean section | 8 (4) |
Place of delivery |
|
Home delivery by traditional birth attendant | 30 (15) |
Hospital delivery by midwife | 120 (60) |
Hospital delivery by doctor | 50 (25) |
Prolonged labor | 136 (68) |
Presentation |
|
Cephalic | 144 (72) |
Breech | 40 (20) |
Transverse | 16 (8) |
Shoulder dystocia | 136 (68) |
Maternal diabetes | 40 (20) |
Maternal age (years)a | 27.5 (6.8) |
Parity of mother |
|
Primipara | 156 (78) |
Multipara | 156 (78) |
aMean (standard deviation), all others are n (%) | |
Mean escape latency of acquisition day | Groups | ||||
NC | SC | ColC | Pre-SwE Exp | Post-SwE Exp | |
Days |
|
|
|
|
|
1st | 26.2 (2.3) | 30.6 (2.4) | 60.0 (0.0)b | 43.2 (1.8)b | 43.8 (1.6)b |
2nd | 22.6 (1.0) | 25.4 (0.6) | 58.9 (0.5)b | 38.6 (2.0)b | 40.5 (1.2)b |
3rd | 14.5 (1.8) | 18.9 (0.4) | 56.5 (1.2)b | 34.2 (1.9)b | 33.8 (1.0)b |
4th | 13.1 (1.7) | 17.5 (0.8) | 53.9 (0.7)b | 35.0 (1.6)b | 34.9 (1.6)b |
5th | 13.0 (1.2) | 15.9 (0.7) | 51.7 (2.0)b | 25.9 (0.7)b | 27.7 (0.9)b |
6th | 12.2 (1.0) | 13.3 (0.4) | 49.5 (2.0)b | 16.8 (1.1)b | 16.8 (0.8)b |
Average of acquisition days | |||||
5th and 6th | 12.6 (0.2) | 14.6 (0.8) | 50.6 (0.7)b | 20.4 (2.1)a | 22.4 (3.2)a |
NC indicates normal control; SC, Sham control; ColC, colchicine control; SwE, swimming exercise exposure. aP <0.05; bP <0.01. | |||||
Categories | Number (%) |
Sex |
|
Male | 36 (60.0) |
Female | 24 (40.0) |
Age in yearsa | 8.8 (4.2) |
Education |
|
Pre-school | 20 (33.3) |
Elementary school | 24 (40.0) |
Junior high school | 16 (26.7) |
Cancer diagnoses |
|
Acute lymphoblastic leukemia | 33 (55) |
Retinoblastoma | 5 (8.3) |
Acute myeloid leukemia | 4 (6.7) |
Non-Hodgkins lymphoma | 4 (6.7) |
Osteosarcoma | 3 (5) |
Hepatoblastoma | 2 (3.3) |
Lymphoma | 2 (3.3) |
Neuroblastoma | 2 (3.3) |
Medulloblastoma | 1 (1.7) |
Neurofibroma | 1 (1.7) |
Ovarian tumour | 1 (1.7) |
Pancreatic cancer | 1 (1.7) |
Rhabdomyosarcoma | 1 (1.7) |
aMean (standard deviation) | |



Test results | Disease | Sensitivity (%) | Specificity (%) | PPV (%) | NPV (%) | ||
Yes | No | ||||||
Reid’s score ≥ 5 | Positive | 10 | 15 | 37.0 | 94.5 | 40.1 | 93.8 |
Negative | 17 | 258 |
|
|
|
| |
Swede score ≥ 5 | Positive | 20 | 150 | 74.1 | 45.0 | 11.8 | 94.6 |
Negative | 7 | 123 |
|
|
|
| |
Swede score ≥ 8 | Positive | 3 | 21 | 11.1 | 92.3 | 12.5 | 91.3 |
Negative | 24 | 252 |
|
|
|
| |
a High-grade indicates a score of ≥5 in both tests; PPV indicates positive predictive value; NPV, negative predictive value | |||||||
Test | Sensitivity (%) | Specificity (%) | Positive predictive value (%) | Negative predictive value (%) |
Reid’s score ≥ 5 | 37.0 | 94.5 | 40.0 | 93.8 |
Swede score ≥ 5 | 74.1 | 45 | 11.8 | 94.6 |
Swede score ≥ 8 | 11.1 | 92.3 | 12.5 | 91.3 |
Test | Sensitivity (%) | Specificity (%) | Positive predictive value (%) | Negative predictive value (%) |
Reid’s score ≥ 5 | 37.0 | 94.5 | 40.0 | 93.8 |
Swede score ≥ 5 | 74.1 | 45 | 11.8 | 94.6 |
Swede score ≥ 8 | 11.1 | 92.3 | 12.5 | 91.3 |
Narakas classification | Total 200 (100%) | Grade 1 72 (36%) | Grade 2 64 (32%) | Grade 3 50 (25%) | Grade 4 14 (7%) |
Complete recoverya | 107 (54) | 60 (83) | 40 (63) | 7 (14) | - |
Near complete functional recovery but partial deformitya | 22 (11) | 5 (7) | 10 (16) | 6 (12) | 1 (7) |
Partial recovery with gross functional defect and deformity | 31 (16) | 7 (10) | 13 (20) | 10 (20) | 1 (7) |
No significant improvement | 40 (20) | - | 1 (1.5) | 27 (54) | 12 (86) |
aSatisfactory recovery bGrade 1, C5, 6, 7 improvement; Grade 2, C5, 6, 7 improvement; Grade 3, panpalsy C5, 6, 7, 8, 9, Grade 4, panpalsy with Hornon’s syndrome. | |||||
Narakas classification | Total 200 (100%) | Grade-1 72 (36%) | Grade-2 64 (32%) | Grade-3 50 (25%) | Grade-4 14 (7%) |
Complete recoverya | 107 (54) | 60 (83) | 40 (63) | 7 (14) | - |
Near complete functional recovery but partial deformitya | 22 (11) | 5 (7) | 10 (16) | 6 (12) | 1 (7) |
Partial recovery with gross functional defect and deformity | 31 (16) | 7 (10) | 13 (20) | 10 (20) | 1 (7) |
No significant improvement | 40 (20) | - | 1 (1.5) | 27 (54) | 12 (86) |
aSatisfactory recovery bGrade 1, C5, 6, 7 improvement; Grade 2, C5, 6, 7 improvement; Grade 3, panpalsy C5, 6, 7,8,9, Grade 4, panpalsy with Hornon’s syndrome. | |||||
Variables in probe trial day | Groups | ||||
NC | SC | ColC | Pre-SwE Exp | Post-SwE Exp | |
Target crossings | 8.0 (0.3) | 7.3 (0.3) | 1.7 (0.2)a | 6.0 (0.3)a | 5.8 (0.4)a |
Time spent in target | 18.0 (0.4) | 16.2 (0.7) | 5.8 (0.8)a | 15.3 (0.7)a | 15.2 (0.9)a |
NC indicates normal control; SC, Sham control; ColC, colchicine control; SwE, swimming exercise exposure. aP <0.01. | |||||
Pain level | Number (%) | P | ||
Pre | Post 1 | Post 2 | ||
Mean (SD)a pain score | 4.7 (1.9) | 2.7 (1.6) | 0.8 (1.1) | <0.001 |
Pain categories | ||||
No pain (0) | - | 1 (1.7) | 31 (51.7) | <0.001 |
Mild pain (1-3) | 15 (25.0) | 43 (70.0) | 27 (45.0) | |
Moderete pain (4-6) | 37 (61.7) | 15 (25.0) | 2 (3.3) | |
Severe pain (7-10) | 8 (13.3) | 2 (3.3) | - | |
aPain scores according to the visual analogue scale ranging from 0 to 10; SD indicates standard deviation | ||||
Surgeries | Number (%) | Satisfactory outcomes n (%) |
Primary surgery (n=24) |
|
|
Upper plexus | 6 (25) | 5 (83) |
Pan-palsy | 18 (75) | 6 (33) |
All | 24 (100) | 11 (46) |
Secondary Surgery (n=26) |
|
|
Shoulder deformity | 15 (58) | 13 (87) |
Wrist and forearm deformity | 11 (42) | 6 (54) |
All | 26 (100) | 19 (73) |
Primary and secondary surgery | 50 (100) | 30 (60) |
Mallet score 14 to 25 or Raimondi score 2-3 or Medical Research grading >3 to 5. | ||
Narakas classification | Total 200 (100%) | Grade-1 72 (36%) | Grade-2 64 (32%) | Grade-3 50 (25%) | Grade-4 14 (7%) |
Complete recoverya | 107 (54) | 60 (83) | 40 (63) | 7 (14) | - |
Near complete functional recovery but partial deformitya | 22 (11) | 5 (7) | 10 (16) | 6 (12) | 1 (7) |
Partial recovery with gross functional defect and deformity | 31 (16) | 7 (10) | 13 (20) | 10 (20) | 1 (7) |
No significant improvement | 40 (20) | - | 1 (1.5) | 27 (54) | 12 (86) |
aSatisfactory recovery bGrade 1, C5, 6, 7 improvement; Grade 2, C5, 6, 7 improvement; Grade 3, panpalsy C5, 6, 7,8,9, Grade 4, panpalsy with Hornon’s syndrome. | |||||
Trials | Groups | ||||
NC | SC | ColC | Pre-SwE Exp | Post-SwE Exp | |
1 | 20.8 (0.6) | 22.1 (1.8) | 41.1 (1.3)b | 31.9 (1.9)b | 32.9 (1.8)a, b |
2 | 10.9 (0.6) | 14.9 (1.7) | 37.4 (1.1)b | 24.9 (2.0)b | 26.8 (2.5)b |
3 | 8.4 (0.5) | 9.9 (2.0) | 32.8 (1.2)b | 22.0 (1.4)b | 21.0 (1.4)b |
4 | 7.8 (0.5) | 10.4 (1.3) | 27.6(1.1)b | 12.8 (1.2)b | 13.0 (1.4)b |
Savings (%)c | 47.7 (3.0) | 33.0 (3.0) | 10.0 (0.9)b | 23.6 (2.7)b | 18.9 (5.3)b |
NC indicates normal control; SC, Sham control; ColC, colchicine control; SwE, swimming exercise exposure. aP <0.05; bP <0.01. cThe difference in latency scores between trials 1 and 2, expressed as the percentage of savings increased from trial 1 to trial 2 | |||||


Table 3 Toxicity profiles of the tested compounds and reference drug tecovirimat
Property | Tecovirimat | Tipranavir | Remdesivir | Fluocinolone | Molnupiravir | Famciclovir | Acyclovir | Cidofovir | Brincidofovir |
Toxicity class | 5 | 3 | 3 | 5 | 5 | 4 | 5 | 4 | 6 |
Carcinogenicity | Active | Moderate | Moderate | Inactive | Inactive | Active | Active | Moderate | Moderate |
Hepatotoxicity | Active | Active | Active | Inactive | Inactive | Inactive | Inactive | Inactive | Inactive |
Cytotoxicity | Moderate | Moderate | Moderate | Inactive | Inactive | Inactive | Inactive | Inactive | Inactive |
Immunotoxicity | Inactive | Moderate | Moderate | Active | Active | Inactive | Inactive | Inactive | Moderate |
Mutagenicity | Moderate | Inactive | Inactive | Moderate | Moderate | Moderate | Moderate | Moderate | Moderate |
The BOILED-Egg model obtained from SwissADME predicted the gastrointestinal absorption and blood–brain barrier (BBB) permeability of the studied compounds. The yellow region represented that molecules could penetrate the BBB. Conversely, the white region corresponded to the compounds with high probability of passive gastrointestinal absorption. The distribution of the tested antivirals indicated that only tecovirimat fell within the BBB-permeable region. Meanwhile, the other compounds remained outside the BBB zone but still demonstrated acceptable absorption characteristics (Figure 1).
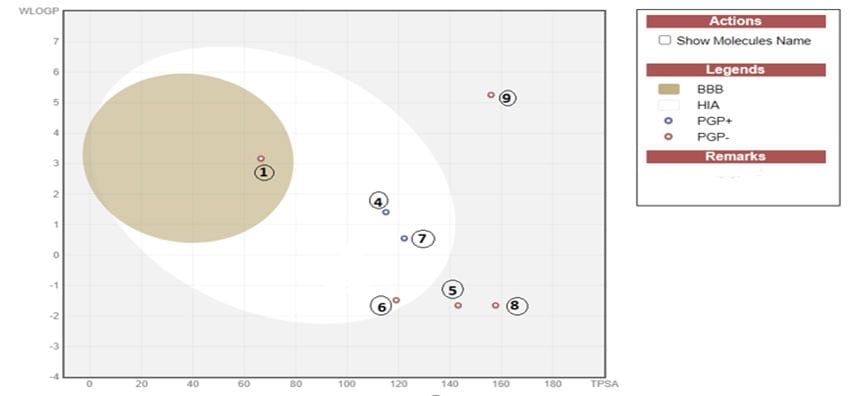
Figure 1 Boiled egg model for compounds (1) Tecovirimat, (2) Tipranavir, (3) Remdesivir, (4) Fluocinolone, (5) Molnupiravir, (6) Acyclovir, (7) Famciclovir, (8) Cidofovir, and (9) Brincidofovir showed blood brain barrier permeability (BBB), P-glycoprotein (P-gp), human intestinal absorption (HIA) of each drug. The yellow zone indicates BBB permeability; blue and red dots indicate P-gp substrate positivity and negativity, respectively.
The bioavailability radar plots summarized six physicochemical parameters that may influence oral drug-likeness. They were lipophilicity, molecular size, polarity, solubility, flexibility, and saturation. The shaded region represented the optimal physicochemical space for oral bioavailability. Here, several tested ligands demonstrated physicochemical profiles comparable to the reference drug tecovirimat which suggested acceptable drug-likeness properties (Figure 2).
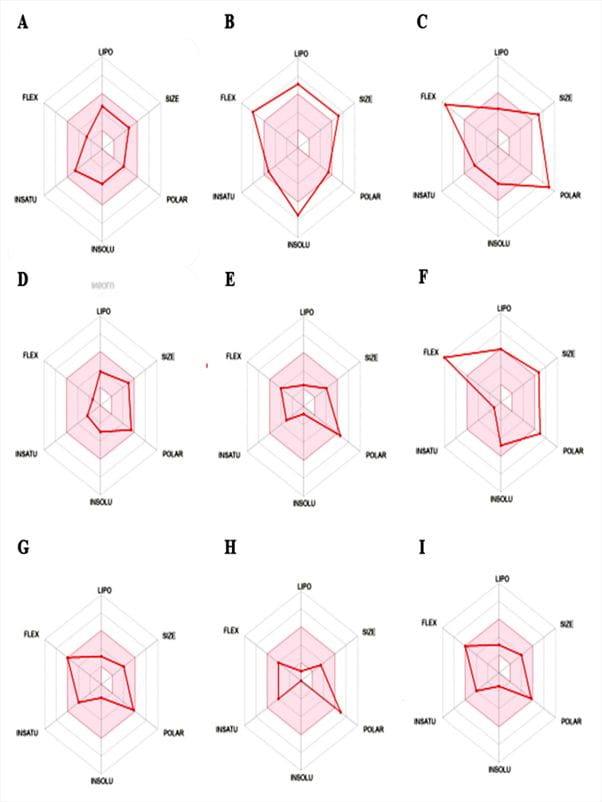
Figure 2 The bioavailability radars of (A) Tecovirimat, (B) Tipranavir, (C) Remdesivir, (D) Fluocinolone, (E) Molnupiravir, (F) Famciclovir, (G) Acyclovir, (H) Cidofovir, and (I) Brincidofovir retrieved from SwissADME. The bioavailability radar illustrates key physicochemical properties influencing oral drug-likeness, including solubility (INSOLU: −6 to 0; good gastrointestinal dissolution), SIZE (150–500 g/mol; suitable permeability and binding), lipophilicity (LIPO: −0.7 to 5.0; better permeability and hydrophobicity balance), flexibility (FLEX: 0–9 rotatable bonds; molecular flexibility for absorption), insaturation (INSATU: Csp³ fraction 0.25–1; structural stability), and polarity (POLAR: TPSA 20–130 Ų; suitable polarity for intestinal absorption). The light reddish region represents the optimal physicochemical space for oral bioavailability; compounds fitting within this area are predicted to possess favorable absorption and permeability profiles.
Lesion-size | Histopathology report | Total | |||||
CIN1 | CIN2 | CIN3 | ICC | CC | SM | ||
0–5 mm | 73 | 0 | 0 | 0 | 5 | 5 | 83 |
6–15 mm | 119 | 18 | 1 | 4 | 0 | 0 | 142 |
>15 mm | 1 | 8 | 31 | 23 | 12 | 0 | 75 |
Total | 193 | 26 | 32 | 27 | 17 | 5 | 300 |
CIN indicates cervical intraepithelial neoplasia; ICC, invasive cervical cancer; CC, chronic cervicitis; SM, squamous metaplasia | |||||||
| Histopathology report | Total | ||||||
CIN1 | CIN2 | CIN3 | ICC | CC | SM | |||
Lesion -Size | 0-5 mm | 73 | 0 | 0 | 0 | 5 | 5 | 83 |
6-15 mm | 119 | 18 | 1 | 4 | 0 | 0 | 142 | |
>15 mm | 1 | 8 | 31 | 23 | 12 | 0 | 75 | |
Total | 193 | 26 | 32 | 27 | 17 | 5 | 300 | |
CIN indicates Cervical intraepithelial neoplasia; ICC, Invasive cervical cancer; CC, Chronic cervicitis; SM, Squamous metaplasia | ||||||||
Group | Didactic posttest marks (%) | Flipped posttest marks (%) | Difference in marks (mean improvement) | P |
<50% | 63.2 (9.4) | 82.2 (10.8) | 19.0 | <0.001 |
≥50% | 72.4 (14.9) | 84.2 ( 10.3) | 11.8 | <0.001 |
Data presented as mean (standard deviation) | ||||





Discussion
This study identified tipranavir, remdesivir, fluocinolone, and molnupiravir as promising poxin inhibitors through molecular docking. Tipranavir exhibited the strongest binding affinity (−9.6 kcal/mol), forming multiple interactions with active site residues. Remdesivir and fluocinolone also showed favorable affinities, while molnupiravir demonstrated binding comparable to tecovirimat [30, 31].
These variables indicate how effectively compounds are absorbed, distributed, metabolized, and excreted in the human body [32]. The potential for oral bioavailability of all the compounds under consideration was demonstrated by their maintenance of a moderate score to the Ghose rule with fewer violations and a virtually good score to Lipinski's rule of five with fewer violations. Specifically, the correlation between TPSA and blood-brain barrier permeability had been examined; a TPSA of less than 90 Å2 was linked to higher permeability, while a TPSA of more than 140 Å2 was linked to lower permeability [33, 34]. Standard drug tecovirimat, with a TPSA of 66.48 Ų, falls within the range associated with favorable oral bioavailability and moderate BBB permeability. This aligned with its clinical use as an oral treatment for smallpox and monkeypox [35]. Even if, Tipranavir, a protease inhibitor used in human immunodeficiency virus (HIV) treatment, had a TPSA of 113.97 Ų. This value suggested a larger molecular size and potentially reduced BBB penetration, which was nearly aligned with the standard drug, tecovirimat. Remdesivir, with a TPSA of 213.36 Ų, exhibited a significantly larger molecular size, indicating limited BBB permeability. This property was consistent with its intravenous administration route, as it was primarily used for treating coronavirus disease (COVID-19) in hospitalized patients [36]. The lipophilicity and solubility profiles of the tested antiviral compounds indicated their suitability for oral administration, as all MLogP values were below 5, satisfying Lipinski’s rule. Tipranavir showed the highest lipophilicity and lowest water solubility, while cidofovir was highly soluble but had low lipophilicity. Most compounds exhibited favorable bioavailability and gastro-intestinal (GI) absorption, although some, including tipranavir and remdesivir, were P-glycoprotein substrates, which may affect their intestinal absorption [27].
The toxicity assessment of the tested antiviral compounds showed a wide range of LD50 values, with brincidofovir (5010 mg/kg) and acyclovir (5000 mg/kg) being the least toxic, while tipranavir (300 mg/kg) exhibited the highest toxicity. Most compounds fell within moderate toxicity classes (3–5), indicating acceptable safety margins at therapeutic doses. The BOILED-Egg model suggested that most antivirals may have good gastrointestinal absorption, while only tecovirimat showed potential blood–brain barrier permeability. Limited BBB penetration for the remaining compounds may reduce possible central nervous system effects while maintaining systemic activity. The bioavailability radar analysis also indicated that several ligands possess physicochemical properties comparable to tecovirimat that showed acceptable oral drug-likeness and pharmacokinetic suitability. Similar ADME prediction approaches are widely used in early drug discovery to evaluate pharmacokinetic feasibility of potential drug candidates [27].
While our findings were encouraging, they were limited by the static nature of docking. The study lacked molecular dynamics simulations to assess binding stability, and the toxicity predictions were computational rather than experimental. Future work should include free energy calculations (Molecular Mechanics/Generalized Born Surface Area, MM-GBSA) and in vitro-in vivo validation to confirm inhibitory potential.
Limitations
This study is based exclusively on in-silico analyses, which have inherent limitations: lack of protein flexibility, absence of dynamic binding validation, and predictive rather than experimental toxicity profiles. Future work should include molecular dynamics simulations, experimental binding assays, and in vitro validation to confirm inhibitory potential.
Conclusion
Our study identified tipranavir, remdesivir, fluocinolone, and molnupiravir as strong potential binders to the monkeypox poxin protein. Their safety and drug-like properties were favorable in our computational analysis. Nonetheless, these predictions require thorough experimental verification. Future essential steps include detailed molecular dynamics simulations, binding assays, and cellular studies to confirm their true antiviral effectiveness against monkeypox infection.



